Sticking sugar to protein
Researchers clarified the 3D structure of an important enzyme that links sugar to proteins – a breakthrough
Whenever cells receive signals, interact with other cells or identify viruses and bacteria, the process involves not only proteins but also sugar chains attached to their surface. The subject of relatively little attention until now, these structures differ widely in composition and branching, and seem to help ensure that proteins fold correctly and are able to perform their specific tasks.
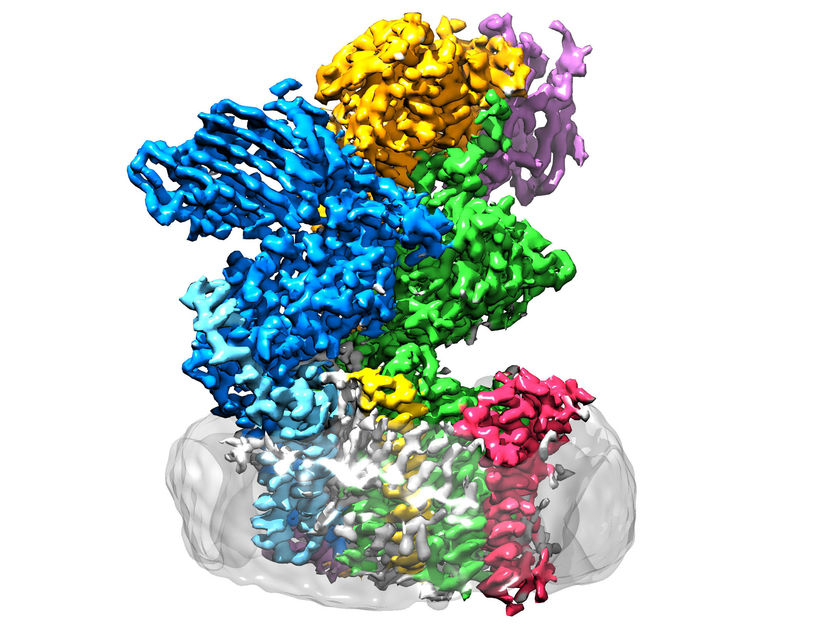
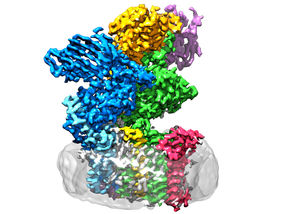
The enzyme oligosaccharyltransferase is located in the membrane (grey) and consists of eight subunits.
ETH Zürich / Julia Kowal
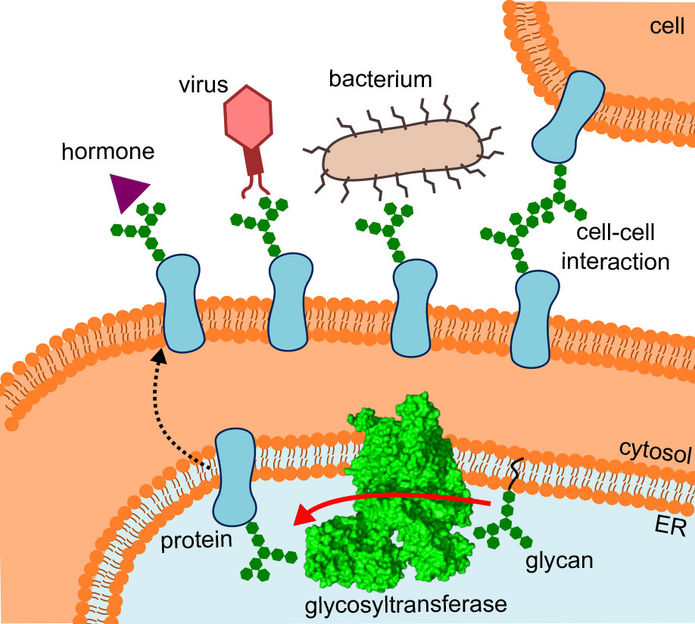
The diagram shows where the glycosyltransferase is located, the pathway the altered proteins take and the interactions they undergo.
R.Wild / ETH Zürich
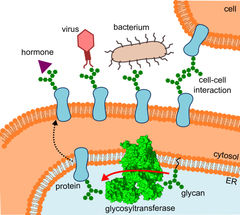
Several observations demonstrate the importance of the sugary appendages: identical proteins with different sugar chains have different functions, diseased and healthy cells have differing sugar compositions on their surfaces, and drugs to which they are attached are apparently tolerated better.
How does the sugar get on to the protein?
Known as glycans, these substances are currently the subject of intensive research. After genomics and proteomics, there is now also glycomics. The emerging research field of glycobiology deals with biosynthesis, structure and the diverse functions of the sugary appendages. One key question is how the diverse set of sugar molecules reaches the proteins in the first place.
ETH researchers in the groups led by Kaspar Locher of the Institute of Molecular Biology and Biophysics and Markus Aebi of the Institute of Microbiology have now taken a decisive step forwards in this area: they have determined the three-dimensional structure of oligosaccharyltransferase (OST) in yeast. “This is the enzyme that connects proteins to sugar trees,” explains Rebekka Wild, one of the three lead authors of the report in Science in which the ETH researchers present their findings.
Made possible thanks to cryo-electron microscopy
Determining the structure of OST was not easy: to start with, Jilliane Eyring, the third lead author of the report, modified the yeast cells so that the enzyme could be targeted and purified. Wild first had to extract the enzyme, which is embedded in a membrane in the cell, from large quantities of these yeast cells and then purify it in a laborious procedure. “Nine litres of yeast produced about 0.2 milligrams of enzyme,” she says. The OST molecules were applied to a small grid, flash-frozen as individual, separate particles and imaged using a high-resolution cryo-electron microscope.
This gave rise to thousands of images showing the enzyme complex from every possible viewing angle. From these images, Julia Kowal, the second lead author, generated the three-dimensional structure of OST. It took six weeks to perform the calculations on the computer cluster. The result was an electron density map which shows the enzyme as an“electron cloud”.
An opportune moment
“We seized on a lucky moment,” Wild explains. They were able to benefit from the “resolution revolution” currently taking place in cryo-electron microscopy and make use of a state-of-the-art device with atomic resolution. The development of this technology was awarded the Nobel Prize in Chemistry in 2017.
In order to interpret the electron microscopy data, Wild had to fit the amino acid sequence of OST “manually” into the small clouds in the three-dimensional map. Thus, the researcher was able to depict the 3D structure of OST in full detail – a breakthrough that was met with delight among researchers after it was published online.
Missing piece of the puzzle
OST is a membrane protein complex made up of eight sub-units, which identify, for example, the sugar or protein substrates or stabilise the catalytic unit. The latter is the sub-unit of the enzyme that harbours the active centre, where proteins and sugars are brought together and fused. “We had expected the catalytic unit to be in the middle of the enzyme,” Wild explains. “Surprisingly, however, it’s on the outside – and the shape is reminiscent of a wide-open mouth.”
What had initially confused the researchers suddenly made sense, as they fitted the structure into the location where OST actually operates: specifically, the enzyme is embedded in vivo in the membrane of the endoplasmic reticulum (ER). This is the part of the cell where proteins are produced, folded, monitored and modified.
Here, the OST’s direct neighbour is a tunnel protein – that much was already known. This funnels the fledgling proteins into the interior of the endoplasmic reticulum and then directly into the wide-open mouth of the OST, where they receive their tiny sugar trees.
Good model for human enzymes
“Yeast OST is a good model for the processes in humans,” Wild explains. The researchers draw this conclusion from the fact that yeast OST’s active centre, where the proteins are fitted with sugars, closely resembles the corresponding area in bacteria. “This means that the active centre has barely changed over the course of evolution,” the researcher explains, “which means there’s a good chance that it still works in a very similar way in mammals, and therefore also in humans.”