Piecing together the full picture of the genome
Revolutionary new DNA sequencing method enables unparalleled rapid, accurate, and efficient analysis while preserving the full picture of a genome
A team of international researchers from the Friedrich Miescher Laboratory of the Max Planck Society developed a new DNA sequencing method that provides fast and efficient tracking of genetic information. Their new technique called haplotagging now enables scientists to quickly and precisely trace and match genome data. Haplotagging delivers rich genome information and facilitates accurate determination of our genetic make-up. This allows for fast genetic analyses in human populations or reveals previously unsuspected mixing of species in nature.
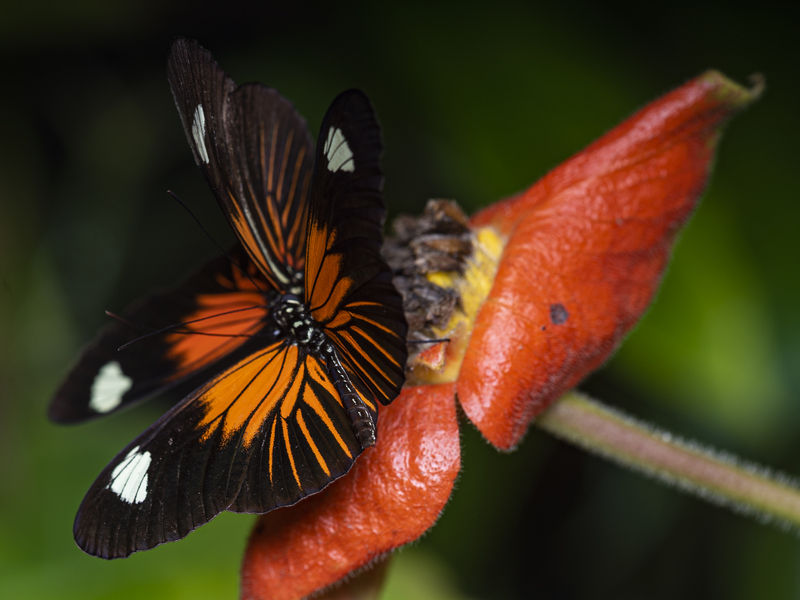

A lowland Heliconius butterfly
Luca Livraghi
Imagine a portrait gallery, filled with the most stunning pictures of people and wildlife. Enchanted, you picked out a favorite and ordered a print. One waits with impatience for the facsimile to be delivered; only to discover that the print comes delivered as shredded pieces, without instructions. Current genome sequencing methods come with similar drawbacks: reading the genome sequence is like reading a book whose pages have been torn out and shredded – the individual words are preserved, but scrambled and fragmented – hardly a satisfying situation and prone to mistakes.
Such is the reality that genome scientists face, having to piece together the big picture—in this case, the DNA sequence of a genome—from many fragmented, individual pieces. The emphasis on sequencing quantity over context means that although we can sequence entire genomes of a person or a pathogen in a single day, they still come in as millions of tiny pieces of “short-reads”, like a genomic jigsaw puzzle. “At best, this complicates things. At worst, we risk missing life-threatening diseases or severe mutations because important information, like scrambled chromosomes, can get lost when we break the genome apart for sequencing. And if we do not manage to put back together what was broken, the big picture is lost,” comments principal investigator Frank Chan, Group Leader at the Friedrich Miescher Laboratory.
Untangling DNA strands like spaghetti
This is where his research group at the Max Planck Campus Tübingen came in, inventing a new DNA sequencing method they called haplotagging. “With this new technique, the configuration of the DNA is marked and preserved, before it is broken up for genome sequencing. This is like numbering jigsaw pieces on the back before breaking them up. Its importance cannot be overstated because it addresses a key short-coming of the current dominant sequencing technology,” Chan explains.
In the lab, this is done by mixing DNA together with microscopic beads. Each bead carries enzymes that pluck out a strand of DNA—much like one can single out an individual noodle from a plate of spaghetti by wrapping it around a fork—and attaches a unique barcode at regular intervals. After breaking the DNA into pieces, this barcode serves to identify which fragments belong together. Together with other practical advantages, the upshot for scientists is that they can now conduct larger genetic studies at lower costs, without compromising quality. “Haplotagging restores some of the original promise of what you should be able to do with sequencing,” Chan adds.
Butterflies flitting with a target on their wings
To showcase the performance of their new sequencing technique, the Tübingen group teamed up with collaborators at the University of Cambridge who have been studying two species of butterflies of the Heliconius genus in Ecuador near the foothills of the Andes. Due to their unique diet, these butterflies can taste awful to their bird predators. And they sport striking, colorful patterns to warn off prying birds. Over time, the two butterfly species have evolved to resemble, or mimic each other: lowland butterflies of both species show striking red rays and one yellow band, and highland butterflies of either species sport two yellow spots set against black wings.
And sure enough, birds learn, and instead pick on rarer, hopefully tastier butterflies that do not display the classic wing patterns found in the two high- and lowland forms. For that reason, hybrid offspring between the two distinct forms may have tragic evolutionary consequences: a mixed up wing pattern is more likely to be targeted by birds. That is why when a new form of hybrid butterflies carrying both the highland double-bar and lowland rays was discovered in the middle of the hybrid zone, it took the researchers utterly by surprise. Using haplotagging to sequence the genome of hundreds of butterflies, Chan and his collaborators were able to decode the genomes and discovered how the spread of a single gene, WntA, from the highlands far into the lowlands was able to give rise to a whole new kind of butterfly, one common enough to keep pesky birds away.
Rapid genetic match-making
The utility of haplotagging is not only limited to studying biodiversity. Chan’s research group also used their technique to help collaborators at the University of Oxford to develop methods for rapid identification of genetic variants. This method, called QUILT, works in a way similar to how text suggestions can finish our sentences for us on our smartphones. By preserving the broader DNA sequence context, haplotagging helps QUILT connect various bits of information and make accurate identification from scattered DNA fragments. “We could show that QUILT performs best with haplotagging data over other sequencing platforms. Haplotagging and QUILT may open up all kinds of possibilities for large population genomic studies,” Chan asserts.
But the impact of haplotagging will also be felt beyond universities and research centers: “This will disrupt the current consumer genetics industry,” Chan predicts. So-called direct-to-consumer companies like 23andMe or Ancestry sell DNA test-kits to everyday consumers, with the promise to discover health risks or finding long-lost relatives. It is now a robust market whose yearly revenue has already surpassed a billion dollars in the U.S. alone. The economic landscape of this sector might drastically change with the arrival of this quick, cost-efficient, and accurate sequencing technique. Ultimately, Chan and his team wants to make life easier for biologists: “Our hope is, in the future, that we spend less time looking down to look at pieces in the DNA puzzle and more time sitting back and admiring the beauty of life and biodiversity.”
Original publication
Meier, J.I.*, Salazar, P.A.*, Kučka, M.*, Davies, R.W., Dréau, A., Aldás, I., Box-Power, O., Nadeau, N., Bridle, J.R., Rolian, C.P., Barton, N.H., McMillan, W.O., Jiggins, C.D.†, Chan, Y.F.†; "Haplotype tagging reveals parallel formation of hybrid races in two butterfly species"; (* co-first authors; † co-last authors); PNAS, 2021.
Davies, R.W, Kučka, M., Su, D., Shi, S., Flanagan, M., Cunniff, C.M., Chan, Y.F.*, Myers, S.*; "Rapid genotype imputation from sequence with reference panels"; (* co-last authors); Nature Genetics, 2021.