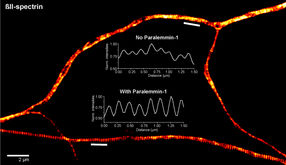
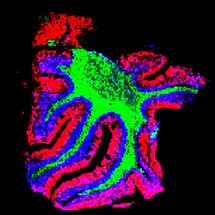
Genetic Fingerprint unmasks Microbial Vandals
For the first time DNA analysis can identify paper-degrading microorganisms. This is made possible by a molecular process developed for fungal infected documents at the University of Vienna with support from the Austrian Science Fund FWF. Fungal species can now be clearly identified by means of a DNA region known as ITS1, making it easier to choose effective countermeasures for conserving historic documents.
It is generally easy enough to say how the ravages of time take their toll on historically valuable papers. Given the right conditions, microorganisms such as fungi can colonise a document and gradually degrade it. However conventional methods for the accurate identification of these fungi are elaborate and imprecise. They require a relatively large amount of sampling material as well as the propagation and subsequent microscopic identification of the fungal sample - a lengthy and error-prone, process. A team led by Dr. Guadalupe Pinar at the University of Vienna Department of Medicinal Chemistry has now developed a process for quickly and unequivocally classifying fungal species on the basis of their DNA.
Dr. Pinar has taken advantage of a special characteristic of the genetic material of many fungal species - a DNA region known as ITS1 which shows enormous differences in the sequencing of DNA base pairs from one strain to another. Outlining the source of these distinguishing features, Dr. Pinar said: "The ITS1 region is often subject to spontaneous mutations. These are harmless as this DNA region doesn¹t have any recognisable function in the fungal genome and plays no direct part in the survivability of a fungal species. But the mutations result in each fungal species¹ having its own typical ITS1 region and therefore a very unmistakable fingerprint." Large amounts of DNA are required to analyse these sequence differences in molecular biological relationships. They could theoretically be obtained by using large amounts of the source material - but that is not an option with historic documents.
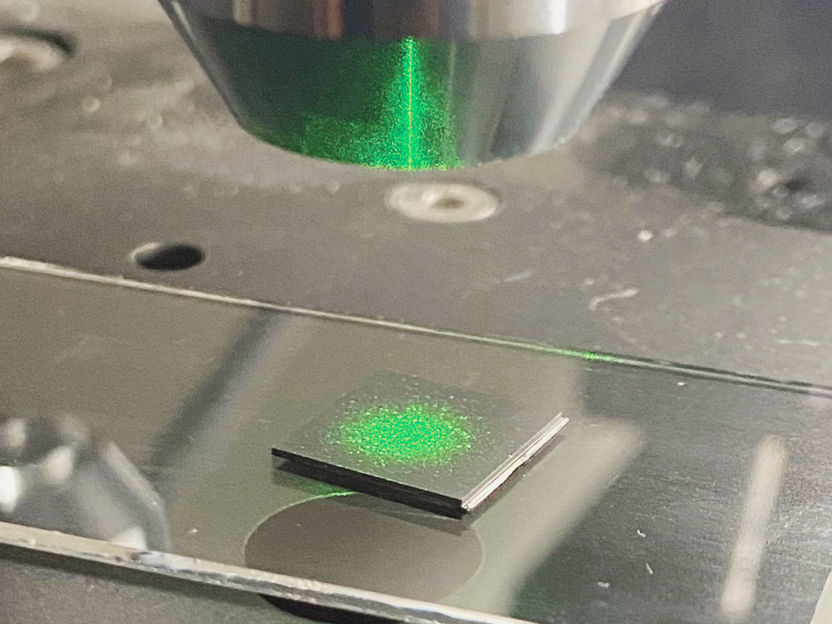
The researchers have now used state-of-the-art methods to clone sufficient quantities of the DNA needed. Once sufficient ITS1 fragments have been cloned the actual DNA analysis can be performed. In a technique known as denaturing gradient gel electrophoresis, the ITS1 fragments are applied to a gel which is subjected to an electrical charge. The ITS1 samples in this field of tension cover different distances depending on the mutations, so each distance is characteristic of a given fungal species. An exchange of even one base pair results in differences which allow the exact fungal species to be identified.
The new method has a further advantage over conventional techniques - even dead fungi can be used as source material. Michaelsen commented: "Fungi become inactive on paper after about 20 years, but the source material for our methods, the DNA, can be isolated from such material as well. This means that samples on which the fungi are inactive but the degradation process is still ongoing can also be investigated using our methods. This is where conventional techniques fall down, as they rely on culturing viable fungi."
Most read news
Organizations
Other news from the department science

Get the analytics and lab tech industry in your inbox
By submitting this form you agree that LUMITOS AG will send you the newsletter(s) selected above by email. Your data will not be passed on to third parties. Your data will be stored and processed in accordance with our data protection regulations. LUMITOS may contact you by email for the purpose of advertising or market and opinion surveys. You can revoke your consent at any time without giving reasons to LUMITOS AG, Ernst-Augustin-Str. 2, 12489 Berlin, Germany or by e-mail at revoke@lumitos.com with effect for the future. In addition, each email contains a link to unsubscribe from the corresponding newsletter.